|
5/16/2023 0 Comments Overlapping dna fragments meaning The strands of the PCR product formed by these extensions act as a pair of oversized primers on the vector fragment. 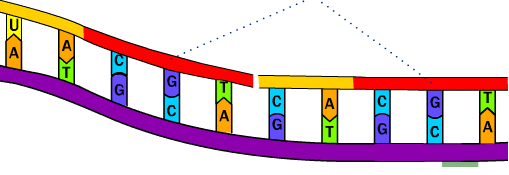
Mode of workingĪ linear with plasmid sequences at both ends insert is created by a PCR reaction. Overlap extension PCR is a straightforward, efficient, and reliable. Recombinases are generally sold as a preoptimized kit by the manufacturer which makes the optimization unavailable for a user to meet their own demands. The requirement of end modifications that cannot be monitored by gel electrophoresis makes TA cloning and LIC less reliable. Any method whose monitoring optimization is easy tends to become the most reliable method. The convenience, price, and efficiency are minor deciding factors. The choice of any cloning method enlisted above is majorly attributed to the reliability of the particular method. Various modification of PCR method has been used for a different cloning purpose. Construction of a recombinant gene encoding a chimeric protein where class I MHC antigen is replaced with class II MHC antigen is discussed here. Such a neat joint is described in this article. Also when no new sequences included the overlap can be designed to make a “neat” joint between two fragments. Base changes incorporated in these regions leads to site-directed mutagenesis. Primers decide the overlap region and they can contain any sequence limited only by the complementary length of the oligomers. This overlap can be extended to form a recombinant product. Then these two fragments are mixed, denatured and reannealed, 3′ end of the top strand anneals with 3′ end of the bottom strand. The ends of these two fragments are modified by mispriming and they share a region of homology. This process is termed as gene Splicing by Overlap Extension (SOE) or gene SOEing. A variant of this method made recombination of different segments from two different genes or “spliced” together by overlap extension. The intrinsic error frequency of this method is sufficiently low, making it practically successful in widespread use. The resultant is a more flexible PCR mutagenesis. The first use of this method is done by introducing mutations into the center of a PCR fragment. This method can make changes at positions close to the restriction sites. the mutation should take place in the primer. This technique is limited by the length of an oligonucleotide from the end of the PCR fragment, i.e. The characteristic of the 5′ end is termed as mispriming it aids in site-directed mutagenesis and in addition of sequences at the end a PCR generated fragment. The extent of this attribute decides the capability of this primers to act as primers for DNA polymerase. #i end primers should be able to meet certain demands such as it should match the sequence of the template gene and at the 5′ ends is capable of including sequences unrelated to the template gene. The result of this process is a DNA segment of defined length this is done by incorporating synthetic oligonucleotide primers into its ends. This process is repeated for multiple rounds this leads to exponential accumulation of the sequence of interest. This strand serves as a template for an extension by a second primer in the opposite orientation. A copy of a DNA strand is formed in this reaction step. 
In PCR process, DNA polymerase is used for extension of the primer. Some related PCR applications are also discussed. In this article, the technique and its uses are discussed briefly. In this PCR based recombination, the reliance on restriction sites is reduced. 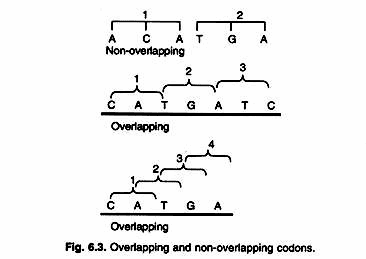
PCR as a synthetic tool can be used for recombining DNA sequences. The dependence on polymerase chain reaction ( PCR) as a fundamental analytical tool for molecular biology tests has increased rapidly. 
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
